Sosnina, E.A., Sosnin, S. & Fedorov, M.V. Improvement of multi-task learning by data enrichment: application for drug discovery. J Comput Aided Mol Des (2023).
DOI
https://doi.org/10.1007/s10822-023-00500-w
Abstract
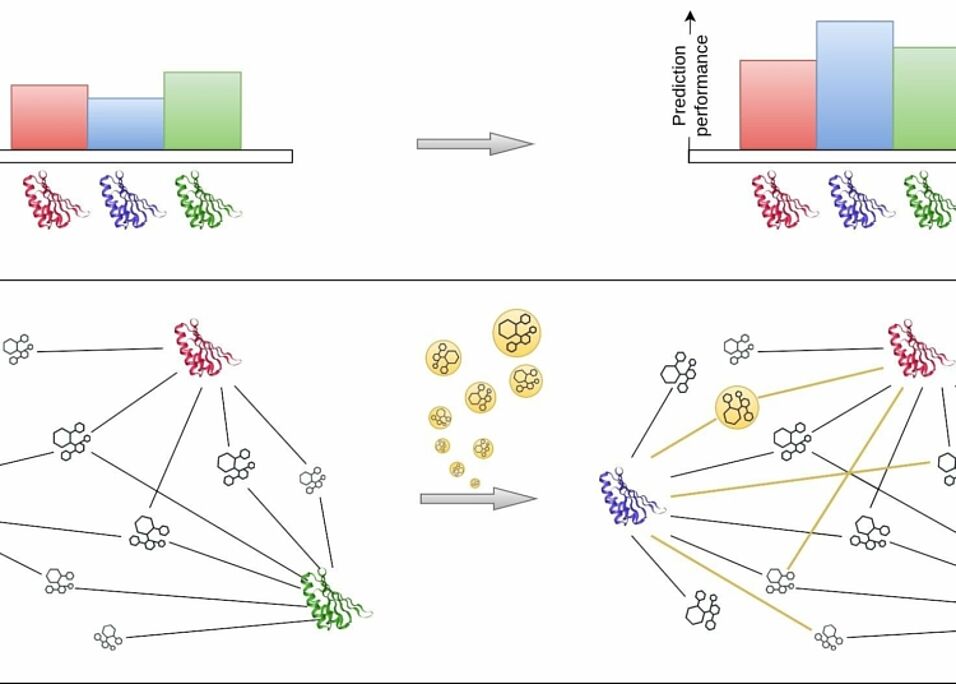
Multi-task learning in deep neural networks has become a topic of growing importance in many research fields, including drug discovery. However, applying multi-task learning poses new challenges in improving prediction performance. This study investigated the potential of training data enrichment to enhance multi-task model prediction quality in drug discovery. The study evaluated four scenarios with varying degrees of information capacity of the training data and applied two types of test data to evaluate prediction performance. We used three datasets: ViralChEMBL, which consisted of binary activities of compounds against viral species, was applied for the classification task; pQSAR(159) and pQSAR(4267), which consisted of bio-activities of compounds and assays from the research of the profile-QSAR method, were applied for regression tasks. We built multi-task models based on the feed-forward DNNs using the PyTorch framework. Our findings showed that training data enrichment could be an effective means of enhancing prediction performance in multi-task learning, but the degree of improvement depends on the quality of the training data. The more unique compounds and targets the training data included, the more new compound-target interactions are required for prediction improvement. Also, we found out that even using multi-task learning, one could not predict the interactions of compounds that are highly dissimilar from those used for model training. The study provides some recommendations for effectively employing multi-task learning in drug discovery to improve prediction accuracy and facilitate the discovery of novel drug candidates.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
Keywords
Multi-task prediction; Deep neural network; Applicability domain; Recommender systems; Data enrichment